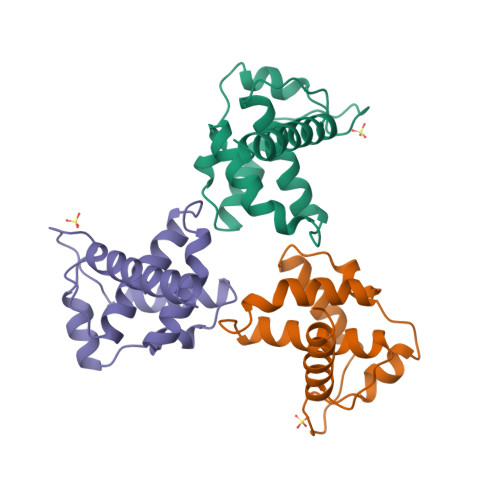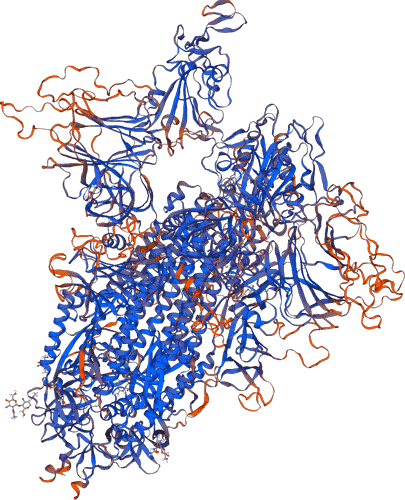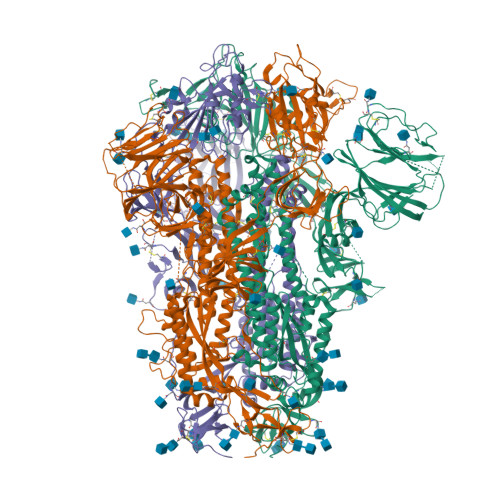
RCSB PDB - 6ZP1: Structure of SARS-CoV-2 Spike Protein Trimer (K986P, V987P, single Arg S1/S2 cleavage site) in Closed State

Metal‐Templated Design of Chemically Switchable Protein Assemblies with High‐Affinity Coordination Sites - Kakkis - 2020 - Angewandte Chemie - Wiley Online Library
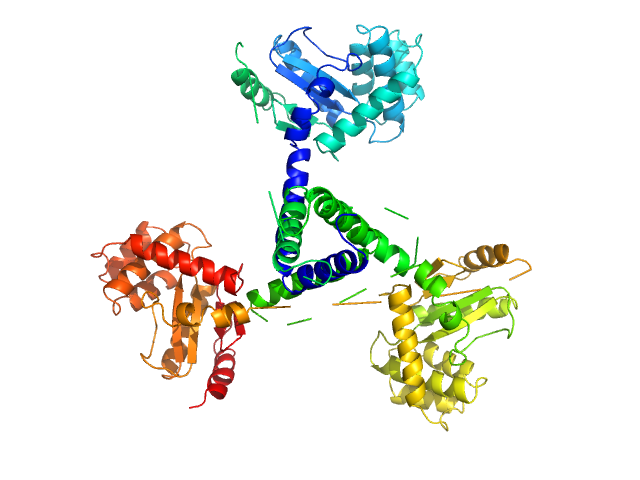
SASDER4 – Wild-type suppressor of copper sensitivity C protein, PmScsC, with concentration series data

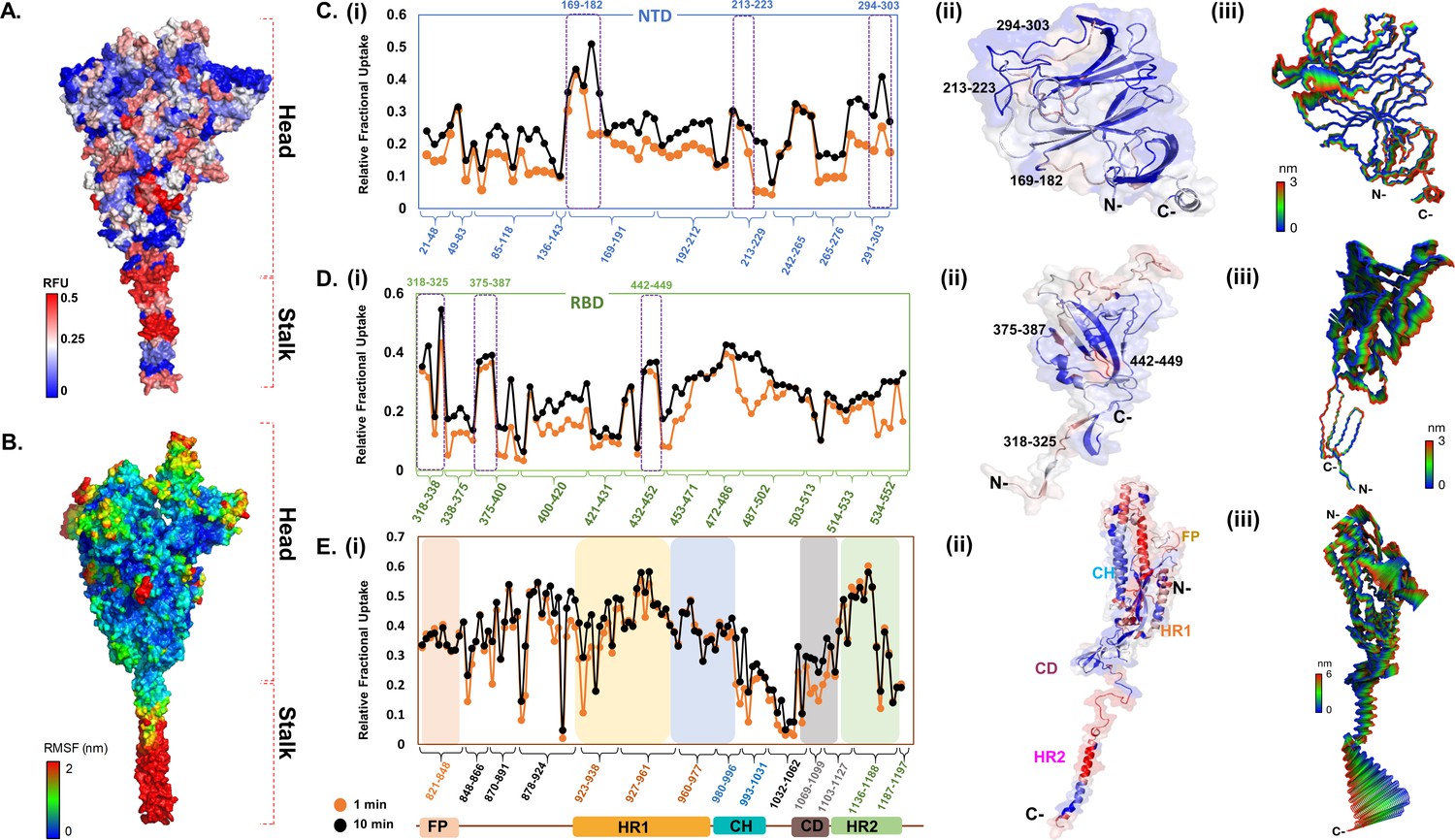
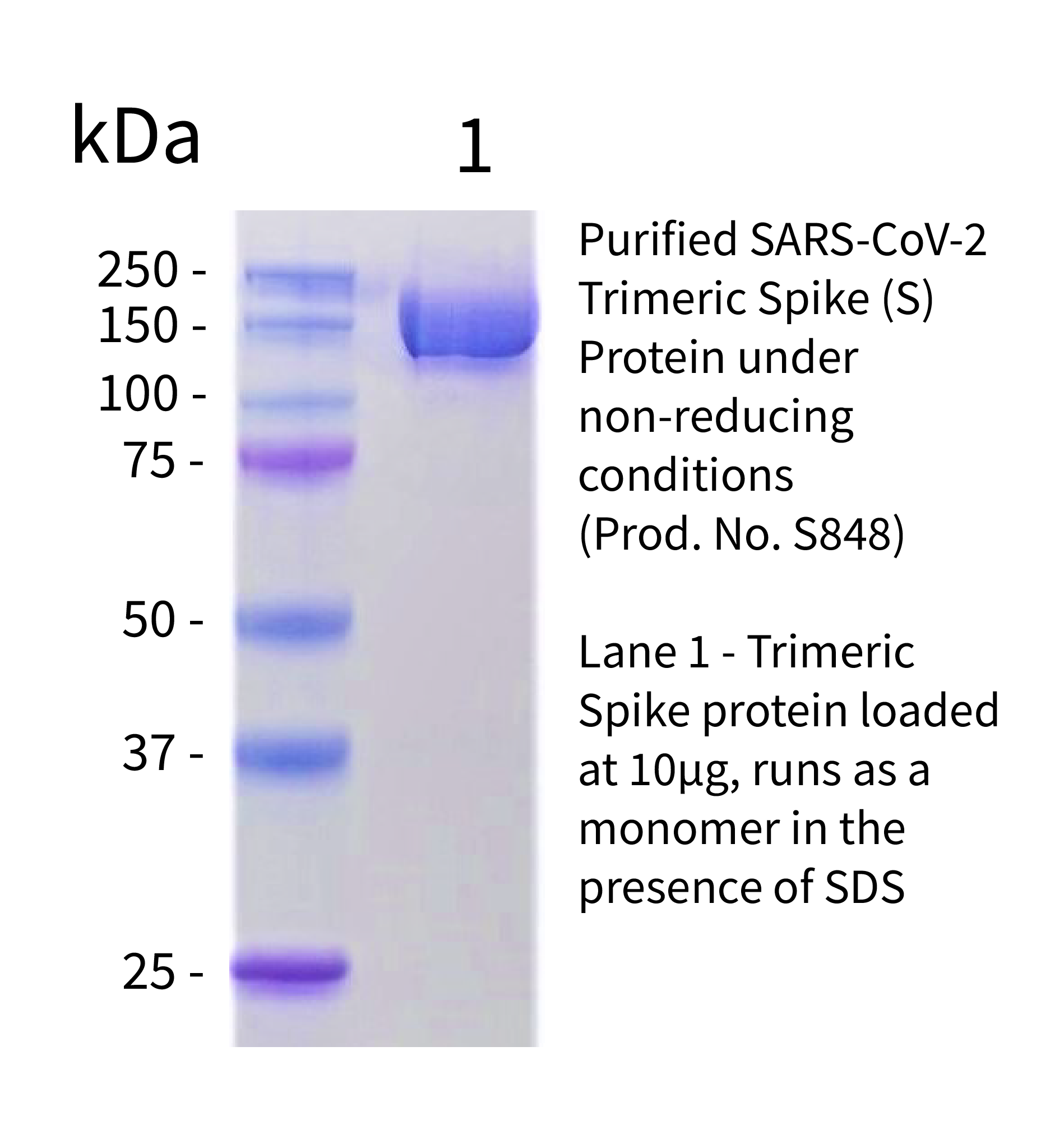
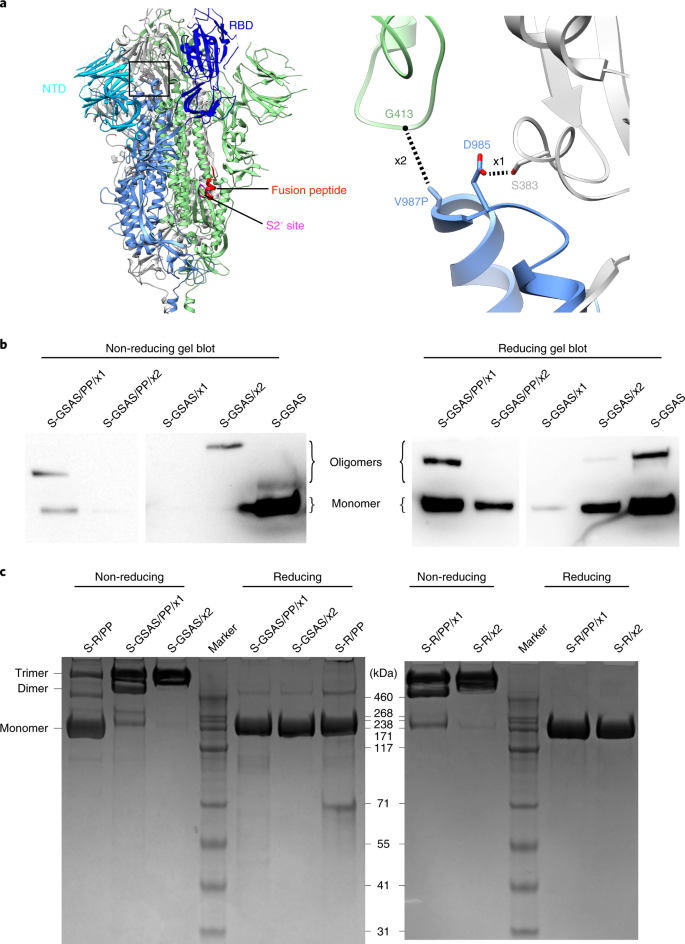
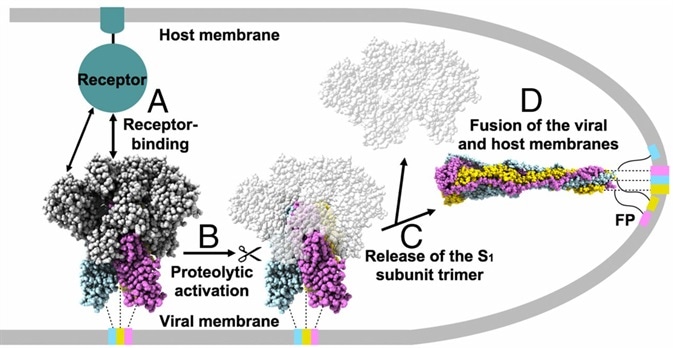
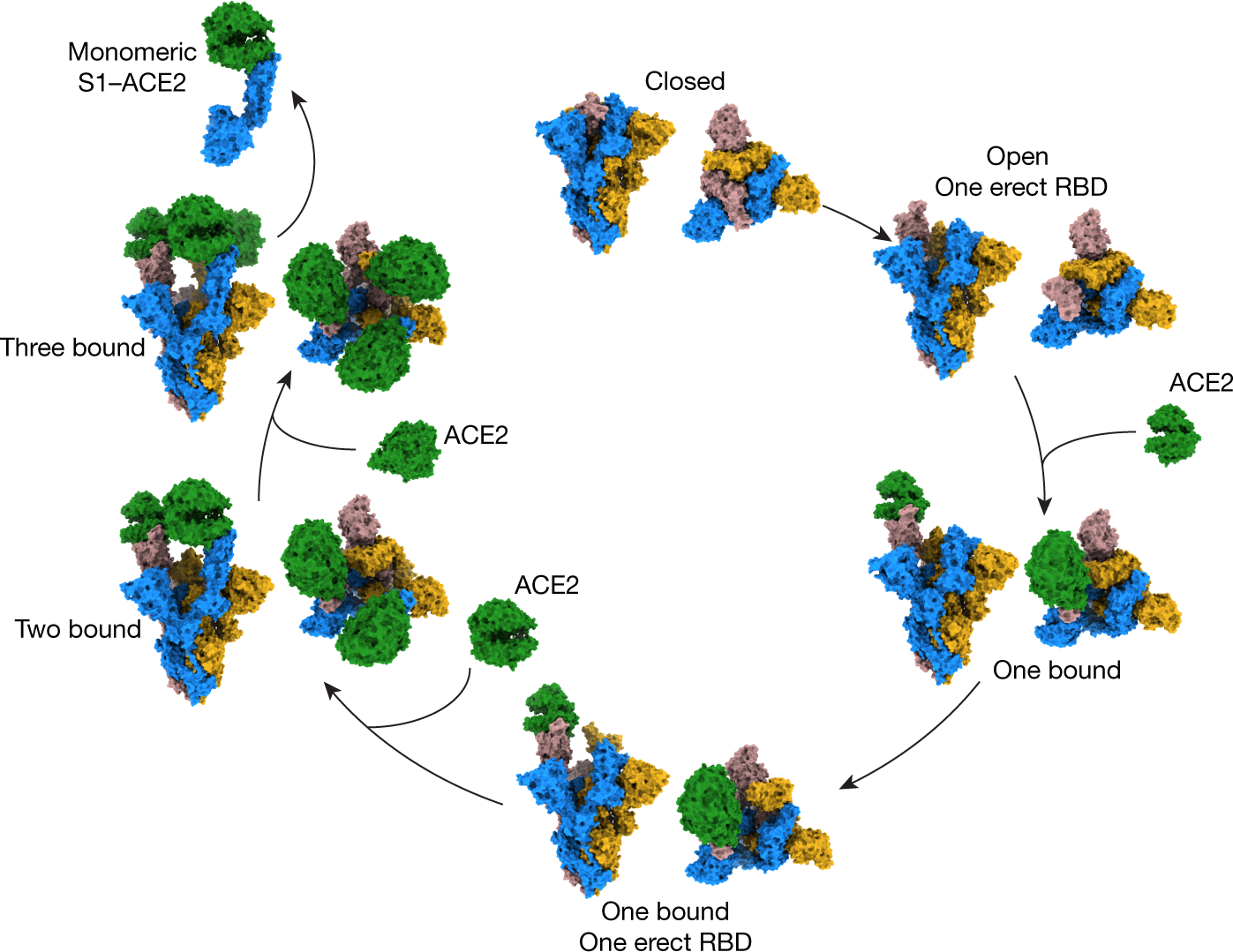
SARS-CoV-2 Spike Protein Stabilized in the Closed State Induces Potent Neutralizing Responses | Journal of Virology
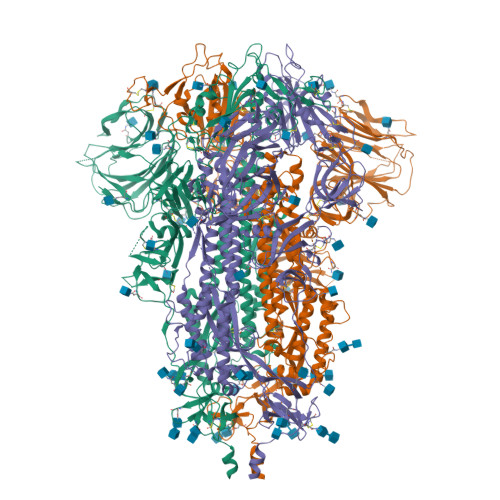
RCSB PDB - 6ZP0: Structure of SARS-CoV-2 Spike Protein Trimer (single Arg S1/S2 cleavage site) in Closed State

Active Participation of Hsp90 in the Biogenesis of the Trimeric Reovirus Cell Attachment Protein ς1* - Journal of Biological Chemistry

Revealing the Activity of Trimeric G-proteins in Live Cells with a Versatile Biosensor Design - ScienceDirect

Disulfide isomerase activity of the dynamic, trimeric Proteus mirabilis ScsC protein is primed by the tandem immunoglobulin-fold domain of ScsB - Journal of Biological Chemistry
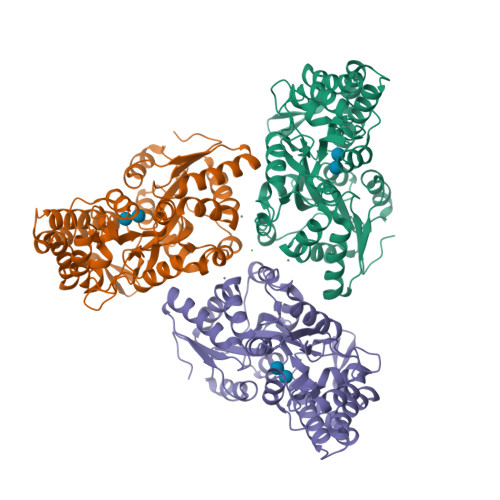
RCSB PDB - 3SEV: Zn-mediated Trimer of Maltose-binding Protein E310H/K314H by Synthetic Symmetrization